Tense DNA will loop neatly
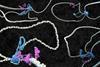
Under a special microscope, the spatial arrangement of DNA turns out to be more complicated than expected.
Want to read more?
Create a free account today!
- Gain access to all our content on chemistry, life sciences and process technology;
- Get our weekly newsletter so you never miss a story.
As a member of the KNCV, KVCV, NBV, or NVBMB you have unlimited access. Log in here.





