Human pluripotent stem cells can become all types of cells, but possess epigenetic mechanisms that block some cellular pathways, a Flemish-Dutch study in Nature Cell Biology reveals.
Naïve human pluripotent stem cells (hPSCs) can in theory differentiate into virtually all possible cell types, just like pre-implantation morula embryos. But the groups of Maarten Dhaenens, Hendrik Marks, Peter Rugg-Gunn and Vincent Pasque have discovered mechanisms that limit the hPSCs.
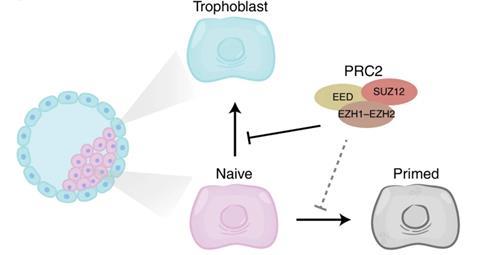
The researchers from Ghent, Nijmegen, Cambridge and Leuven respectively looked specifically at the formation of trophoblast cells from hPSCs. Trophoblast cells normally arise four days after the fusion of a human sperm and egg cell from the morula embryo and form the basis for the placenta. However, trophoblast cells do not normally arise spontaneously from hPSCs, suggesting that formation is actively suppressed. Using a multi-omics approach, they investigated what that barrier might be.
Omics techniques are studies that examine a whole panel of biomolecules’, Maarten Dhaenens (UGent) explains. We used different omics technologies (together ‘multi-omics’, ed.) to study the proteins, DNA and RNA at the earliest possible regulatory stage of the hPSCs. We focused on the epigenetic mechanisms that decide which genes are activated and ultimately translated into proteins. Here, histone modifications and DNA methylations play a very important role.
Our genetic material is more complex than just the DNA double helix. Hendrik Marks (Radboud): ‘DNA is wrapped around histones, and its expression into RNA and other processes is regulated by transcription factors and enzymatic processes. The latter open up the DNA for transcription, depending on whether a transcription factor binds to the gene and can recruit the other enzymes.’
Brake
Multi-omics can be roughly divided into two fields: mass spectrometry (MS) for proteins on the one hand, and sequencing or reading DNA and RNA on the other. ‘In epigenetics these two come together’, says Dhaenens. ’With MS we saw that a certain histone modification occurs much more frequently in stem cells that have chosen a direction than in naive stem cells. We then used sequencing to find out where on the DNA the spindles (coil-like structures of histones around the DNA, ed.) with that modification were located.
The histone modification that the researchers found very specifically in the hPSCs is made by the protein complex PRC2. It is a trimethylation on lysine number 27 of histone 3, abbreviated H3K27me3. ‘Interestingly, we found that H3K27me3 was on genes that are important for placental development,’ adds Marks. Those genes were found to be turned off, thus blocking placental development. ‘After those steps, we tried to inhibit PRC2 and thus H3K27me3 with small molecules. Subsequent RNA sequencing showed that the cells without the H3K27me3 inhibitor were indeed much more efficient in moving towards placental cells.’ The PRC2 protein complex thus acts as a kind of gatekeeper.
Simultaneous
According to the researchers, this fits in well with earlier observations. ‘We also saw high H3K27me3 in pluripotent stem cells from mice’, says Dhaenens. Simultaneously with the current study, another group published a paper on exactly the same subject in the same journal. ‘That is very important to us, a sign that our findings are very repeatable and robust. It gives us the feeling that we are indeed moving in the right direction, it makes us more confident about our work.’
‘We’ve collected great data,’ says Marks. ‘We have benefited greatly from recent developments in embryonic systems. Last year Nicolas Rivron, one of the co-authors working at a research institute in Vienna, published a new method for growing embryo-like structures. ’We were able to use those models powerfully to test our findings, which produced beautiful microscopic images (see below, ed.) that clearly show the role of H3K27me3.’
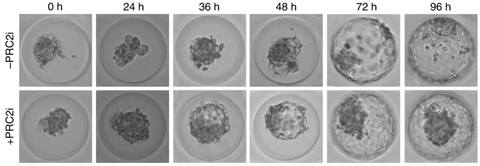
Zijlmans, D.W. et al. (2022) Nat Cell Biol 24, doi.org/10.1038/s41556-022-00932-w











Nog geen opmerkingen