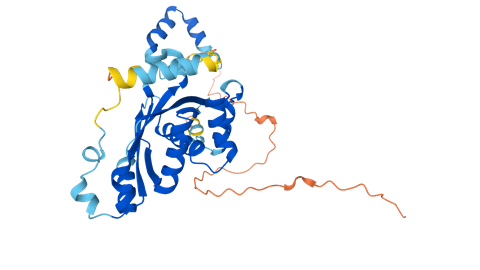
Science and Nature consider the prediction of protein folding with artificial intelligence to be the scientific breakthrough of 2021. We spoke to two experts about the opportunities and limitations of this ‘giant leap forward’.
One day it will be possible to predict the three-dimensional structure of a protein solely on the basis of its amino acid sequence, predicted biochemist and protein structure expert Christian Anfinsen in his acceptance speech for the 1972 Nobel Prize. Almost fifty years later this prophecy is fulfilled: last year, developers managed to predict protein folding almost perfectly using artificial intelligence (AI).

Previously, determining protein structures required labor-intensive analyses in the lab. Everything changed in July 2021, when the Google DeepMind team published their AlphaFold method in Nature. Shortly thereafter, the Baker Laboratory at the University of Washington (Seattle) followed with their similar method RoseTTAfold in Science. Both methods use a deep learning algorithm and can calculate protein structures relatively quickly and easily. The DeepMind team has since applied AlphaFold to several complete genomes, including that of humans. They predicted the structure of almost every protein in the human body and the nearly complete proteomes of twenty other organisms, including the mouse, fruit fly and Baker’s yeast (Saccharomyces cerevisiae).
Giant step
AlphaFold uses fifty years of experimentally obtained structure information on proteins from the Protein Data Bank (PDB) and all available sequence data in various databases. It looks at evolutionary changes by comparing and integrating all sequences with what is known about structures. Combining all the information, AlphaFold learns how an amino acid sequence might fold. While doing so, it uses colors and a matrix to indicate how reliable the predictions are.
‘It’s like having the internet at your disposal versus still having to write the book’
Titia Sixma
‘It works wonderfully!’ says Titia Sixma. ‘You can see what the protein structure of proteins you haven’t measured could be like.’ Sixma is a structural biologist at the Netherlands Cancer Institute (NKI) and a frequent user of AlphaFold. She investigates the mechanism of action of proteins and the signaling processes that regulate the repair of DNA functions. This information is important for, among other things, developing effective cancer drugs.
‘We didn’t expect to see this in our scientific lifetime,’ Sixma says. ‘Structure information is very useful and has often led to Nobel Prizes because it gives so much information. AlphaFold is a giant step forward in understanding.’ In the past, structural biologists had to dabble for years - synthesizing and purifying protein, applying X-ray crystallography with complex, expensive and over-demanded equipment - to visualize a protein structure, now they can ‘just take a look’ based on the amino acid sequence. AlphaFold shows where the domain boundaries are and provides information on activation due to binding at a location other than the active center, called allosteric activation.
Multimers and conformational changes
Some of the specific functions required of the software were already known, namely the part that predicts interactions between amino acids that are close together in the chain. These functions were originally developed to process natural language: it must link groups of words that are close together. This so-called ‘attention network’ tries to take all the input at once and constantly balance everything out. ‘The real problem was interactions between amino acids that are far apart,’ says Anastassis (‘Tassos’) Perrakis, Sixma’s colleague at the NKI and working on the automated building of protein models. ‘That really required fundamental development. They had to invent new mathematics to write the transformer functions that could predict these interactions.’
‘They had to invent new mathematics to write the transformer functions that could predict these interactions’
Tassos Perrakis
Although Sixma and Perrakis are excited about the possibilities, the models cannot answer most of the questions they have as researchers. AlphaFold is limited to proteins and does not include interactions with partner proteins or multimers (dimers, trimers, tetramers). Co-factors, metal ions and other ligands are also not considered. Sixma: ‘Alphafold, for example, builds up hemoglobin without a heme and as a monomer. That does not exist in a realistic environment. The structure is correct in principle, but the heme is missing and it should actually form a tetramer.’ This is now partly solved by Alphafill, a web server created by Perrakis and his research group, which places known ligands into the Alphafold models.
RoseTTAfold did do something with multimers, and successor AlphaFold-Multimer can predict homomeric and heteromeric interactions, albeit with variable accuracy. Perrakis would like to look at even larger multimers. Currently, you can look up complexes up to a certain size, after that a computer’s graphics card can’t handle it. The dataset describing the protein then becomes so large that the graphics card can no longer process that amount and therefore cannot convert it into an image.
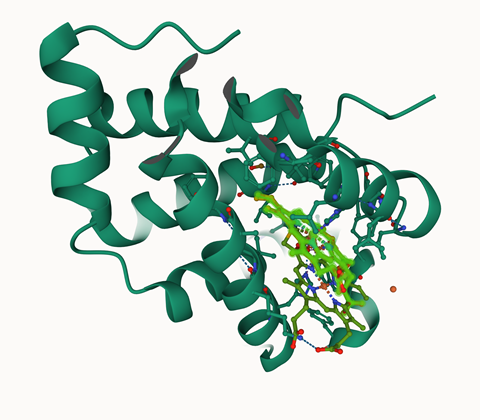
For Sixma, the main shortcoming is that AlphaFold predicts only one state. A real protein undergoes conformational changes due to binding to other proteins, context, and chemical modifications such as phosphorylation and glycosylation; the tool does not show which of the possible conformations it predicts. Sixma actually wants to see how its function changes when the protein undergoes a conformational change. ‘For example, AlphaFold often grabs the active conformation if it is more common in the PDB.’
In addition, AlphaFold is not suitable for predicting the effect of a point mutation. ‘AlphaFold takes information from the evolutionary multisequence alignment. I sometimes see people predict the structure of a mutation in the covid variant omicron at key points. It’s not actually suitable for that,’ Sixma says. By now there are disclaimers in the AlphaFold database. ‘The danger is that the user puts too much value on the prediction.’
Small molecules
AlphaFold also cannot predict RNA or DNA structures yet. Those structures are less complicated than proteins, and Sixma says that function may be added in the future. ‘But the problem with DNA is that we have much less information in the PDB. It’s only a small fraction compared to the known protein structures.’
Another missing function is the binding of small molecules, such as many drugs. Small molecules are more variable and our understanding of that interaction with proteins is really still lacking, according to Sixma. ‘That dream was already there when I was a student. For protein folding, they had a great source of information, but binding of small molecules can still take a long time. AI always needs a lot of examples.’ Perrakis adds: ‘The biggest problem with this is the inaccuracy in placing side chains in the current version of Alphafold. But an update costs millions of euros. It’s only worth it if they can achieve a significant improvement.’
‘You still need structural biology to understand how it really works. We just let the machine do all the boring modeling work’
Titia Sixma
Biologists can now access a huge source of information with little effort, although Sixma says there is also a danger in this. ‘People may think that we don’t need to do any more experiments with proteins. But to understand how it really works, you still need structural biology. We just let the machine do the boring modeling work. But it doesn’t show large complexes, it doesn’t show which specific state does what, and DNA and RNA are missing. That combination is precisely what is interests us.’

Crystallography therefore remains important for the time being, especially for drug development and binding of small molecules. Sixma, Perrakis and their colleagues now mainly use cryo-electron microscopy (cryo-EM) to gain insight into how proteins work. Sixma does expect AlphaFold to be a huge boost to biochemistry. Previously, researchers had to wait one or two years before they could answer a question; now, all they have to do is look at the computer. ‘It’s like having the internet at your disposal versus still having to write the book.’
Pure proteins
Therein also lies immediately the greatest objection. Protein crystallographs produce a lot of pure protein to determine a structure. This is a complicated, labor-intensive task, and it takes new researchers a year to master the necessary techniques. They could then use the purified protein for a variety of other biochemical applications. Now that some of the need is gone, people may not bother.
The consequences of this are as yet incalculable. Sixma: ‘It is not easy to purify a protein properly. But if you don’t, you get contaminated enzymes. Arthur Kornberg (American biochemist who received the Nobel Prize for Physiology and Medicine in 1959 for the mechanisms of the biological synthesis of DNA, ed.) said: ‘Don’t waste clean thinking on dirty protein’. If you’re doing in vitro experiments and biochemistry and want to be quantitative, you have to have those pure proteins. If AlphaFold is going to mean that people don’t make those anymore, that could limit research in some places.’
Structural Biology
Structural biology is a branch of molecular biology, biochemistry and biophysics that studies the spatial structure of biological macromolecules, primarily proteins, RNA, DNA and membranes. Structural biology studies how these macromolecules organize themselves and how changes in their structure affect biochemical function. Biomolecules are too small to see with a light microscope, which is why structural biologists use techniques such as X-ray crystallography, NMR and cryo-EM to determine structures.
Structural biology has provided insight into numerous molecular components and mechanisms that play a role in physiology and disease. It is also an important tool in the search for new drugs.